DebiasedDTA: A Framework for Improving the Generalizability of Drug-Target Affinity Prediction Models
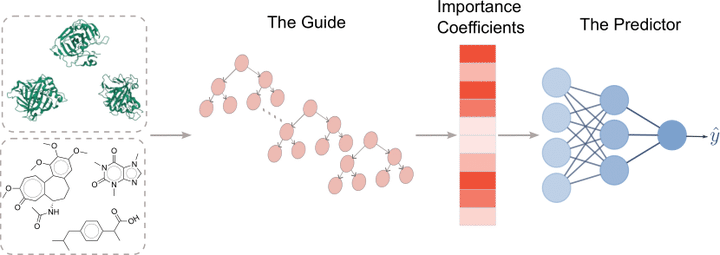
Abstract
Computational models that accurately predict the binding affinity of an input protein-chemical pair can accelerate drug discovery studies. These models are trained on available protein-chemical interaction datasets, which may contain dataset biases that may lead the model to learn dataset-specific patterns, instead of generalizable relationships. As a result, the prediction performance of models drops for previously unseen biomolecules, i.e. the prediction models cannot generalize to biomolecules outside of the dataset. The latest approaches that aim to improve model generalizability either have limited applicability or introduce the risk of degrading prediction performance. Here, we present DebiasedDTA, a novel drug-target affinity (DTA) prediction model training framework that addresses dataset biases to improve the generalizability of affinity prediction models. DebiasedDTA reweights the training samples to mitigate the effect of dataset biases and is applicable to most DTA prediction models. The results suggest that models trained in the DebiasedDTA framework can achieve improved generalizability in predicting the interactions of the previously unseen biomolecules, as well as performance improvements on those previously seen. Extensive experiments with different biomolecule representations, model architectures, and datasets demonstrate that DebiasedDTA can upgrade DTA prediction models irrespective of the biomolecule representation, model architecture, and training dataset. Last but not least, we release DebiasedDTA as an open-source python library to enable other researchers to debias their own predictors and/or develop their own debiasing methods. We believe that this python library will corroborate and foster research to develop more generalizable DTA prediction models.